When talking about getting AI into clinical practice faster, one of the most important factors is algorithm performance. The AI software should be reliable and accurate, boost radiologist performance, help save time and, of course, at the end of the day, improve patient lives. All of this requires algorithms that perform well on the medical data of a wide range of clinical facilities.
How can we assure that AI radiology algorithms are up to this task? As long as there is sufficient data available for training, good performance should be within reach. Nevertheless, there is a lot you can do by organizing the data in a smart way before training. Here we discuss one particular tweak, stratification, that can be used to give algorithms a bump in performance.
The start of training an AI radiology algorithm
First, let’s take a look at the radiology dataset you will need for training. Or actually, the three datasets. Indeed, to develop a well performing AI radiology algorithm, actually three different datasets are needed. Usually these are obtained by starting with one large dataset and simply splitting this in three. All these three datasets play their unique and important role in the algorithm development process.
Start with a training set and a validation set
The algorithm development process starts with initial training of the network. This actually requires two of the three sub-datasets already: the training set and the validation set. Since these datasets collaborate intimately, we will discuss them in one go.
During training, the training dataset is fed to the neural network to create the backbone of the algorithm. The network looks at the examples in the training set and will learn to recognize certain patterns. While training, the validation dataset is used as an independent check of algorithm performance. This means that while the network is learning to recognize image features that make up, for example, a prostate lesion in the training set, you independently verify performance on prostate lesions in the validation set. This allows you to check that the network is not overfitting on the training set (i.e., the network actually starts to recognize each sample in the training set). This performance check with the validation set is done multiple times during training. Furthermore, this test can be used to choose when to stop training and save the final version of the algorithm. Usually, this is done when the performance on the validation set is at its peak. In most cases about 80% of the total dataset is reserved for the training and validation combined. Approximately three quarters (60% of the total) is used or the training set and the remaining quarter (20% of the total) for validation.
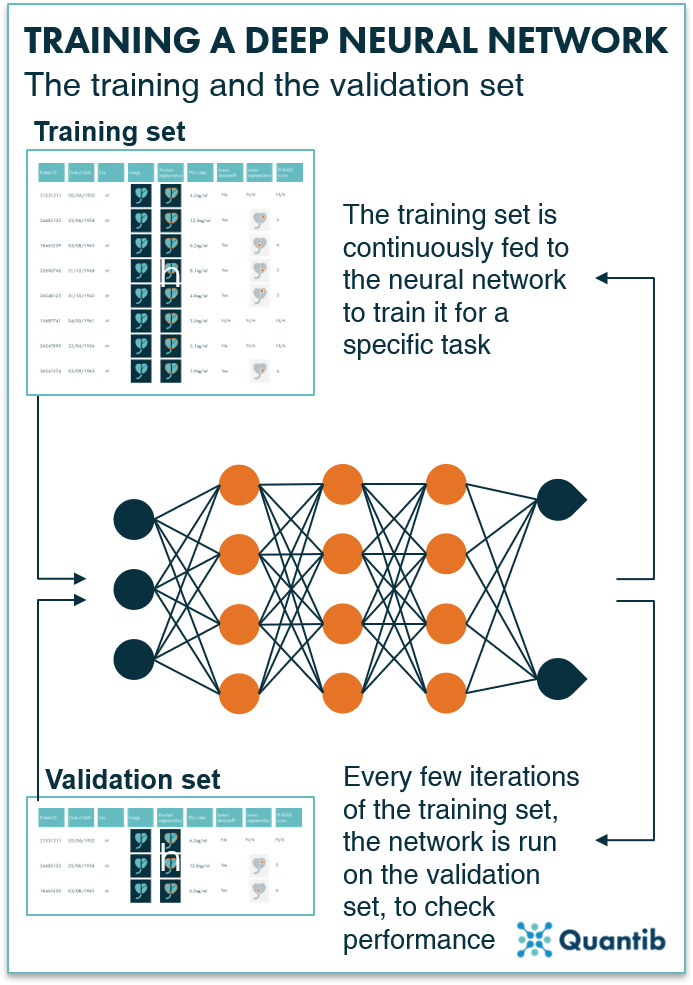
Figure 1: Both the training and the validation sets are used to train the neural network. The training set is used for continuous training and the validation set is used to optimize the parameters further with an independent check.
The final check: applying the test set
The remaining 20% of the complete dataset is meant to serve as the test set. The test set is meant to be a truly independent dataset and should never have been involved in the training of the AI network. In fact, to determine the final performance of the ,,,,asdfdf, e.g. to file for regulatory approval, this dataset should not be used more than once. Reserving this dataset until the very end ensures you that the algorithm is not tweaked in such a way that performance is better on the test set (which would cause it to lose generalizability to data it may encounter in the future). Executing the last test-set-check should optimally resemble the performance in real-world scenarios, so that it is clear the algorithm is up to the task of being fully embedded in clinical practice. It is therefore not surprising that this is the algorithm performance regulatory bodies evaluate when determining whether or not a software product should be cleared for clinical use.
In an ideal world, performance on the validation set and test set would be equal. However, because the validation set is used to finetune the hyperparameters of the algorithm, it is still common to see a small dip in performance when applying your carefully crafted algorithm to the test set, as this is brand new data that the algorithm has not seen before. This should be kept in mind during development, since it may hamper regulatory approval if the dip in performance is too large.
How to boost performance of an AI radiology algorithm
Now that the difference between these data subsets is clear, this knowledge can be used to our advantage to improve algorithm training. Besides keeping track of the ratios between the different sets, it is important to bear several other things in mind when selecting data to create the different datasets. Below we will focus on different ways to split the one radiology dataset you started with into all these different sub-sets.
Approach 1: plain old randomization
The most straightforward approach is randomization. Just make sure that random really means random, so give the dataset a good shake (make sure it is not accidentally sorted by study date, patient ID, etc) before you select the first 60% of cases for training, then 20% for validation, and finally 20% for testing.
However, particularly for small datasets (and mind you that medical datasets are typically pretty small compared to the datasets used in other AI venues, one way to overcome this challenge, is by using augmentation), this is likely to result in unfair distributions. For example, more patients with high PSA values might end up in the training set by accident, while the validation set contains relatively small prostates, and test set has mostly cases with tumors in the peripheral zone, and so on. This introduces bias in the algorithm and will lower performance on the validation and test set (and of course in clinical practice, which is what it is all about eventually). Although the chances of obtaining fair distributions are greater for relatively large datasets, the safe route is going for approach #2, stratification.
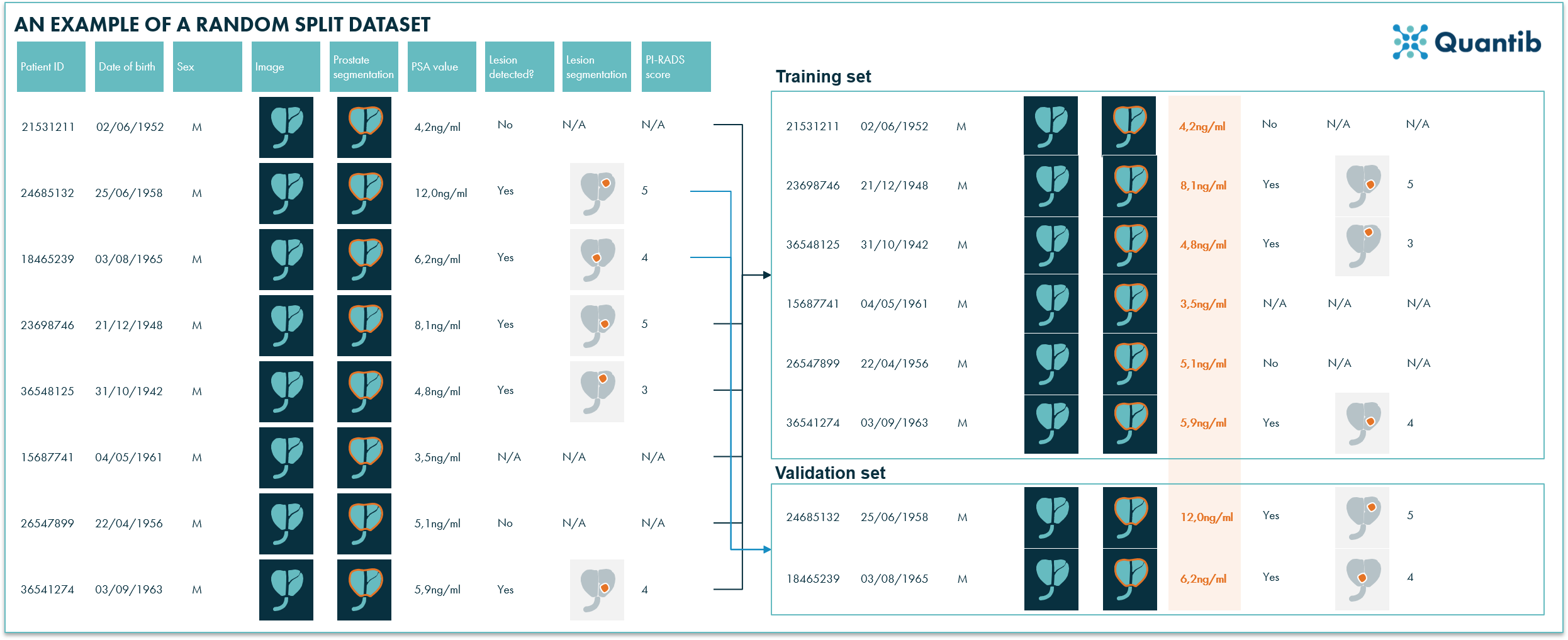
Figure 2: In this example a randomized split resulted in a training set with relatively low PSA values and relatively high PSA values in the validations set. Ideally, a training- and a validation set have a similar distribution of all features, hence randomized splitting is not the best way to distribute the data entries over the different sub-datasets.
Approach 2: stratification
A far better, but trickier to implement approach is stratification. Stratification simply means that you take a good hard look at each and every example in the dataset, and determine which of the various datasets this example should be placed in. If you see that your training set already contains a large number of cases with high prostate volumes, you can try to add some cases with small prostate volumes. In this way a nice distribution of patients with large prostate volumes / high PSA values / low PIRADS scores / etc. can be created over the different sub-sets.
Of course, doing this by hand will take ages. Even for smaller radiology datasets this will be a very time-consuming endeavor. Automation is really the way to go, which means you need a separate stratification algorithm to help you perform this task. This algorithm iterates over all patients in the dataset and calculates for each patient the most suitable data subset to which it should be assigned, depending on which cases are already in this subset.
Some stratification examples
In the end, all subsets should be relatively similar, that is, contain a representative number of patients with large prostates, no lesions, high or low age, etc. The complicating factor is that these different features may or may not be correlated. Older men may have larger prostates on average, but there may be no correlation between age and the number of prostate lesions. When you are iteratively deciding for each case which dataset to assign it to, based on the prostate volume the patient may be most suited for the training set, but based on the number of lesions it might be most suited for the validation set. Add a large number of other stratification parameters in the mix and the problem gets ever more complicated. A stratification algorithm simultaneously keeps track of all these features and optimizes the distribution of all data entries over the different subsets. Visually, this will look something like figure 3.
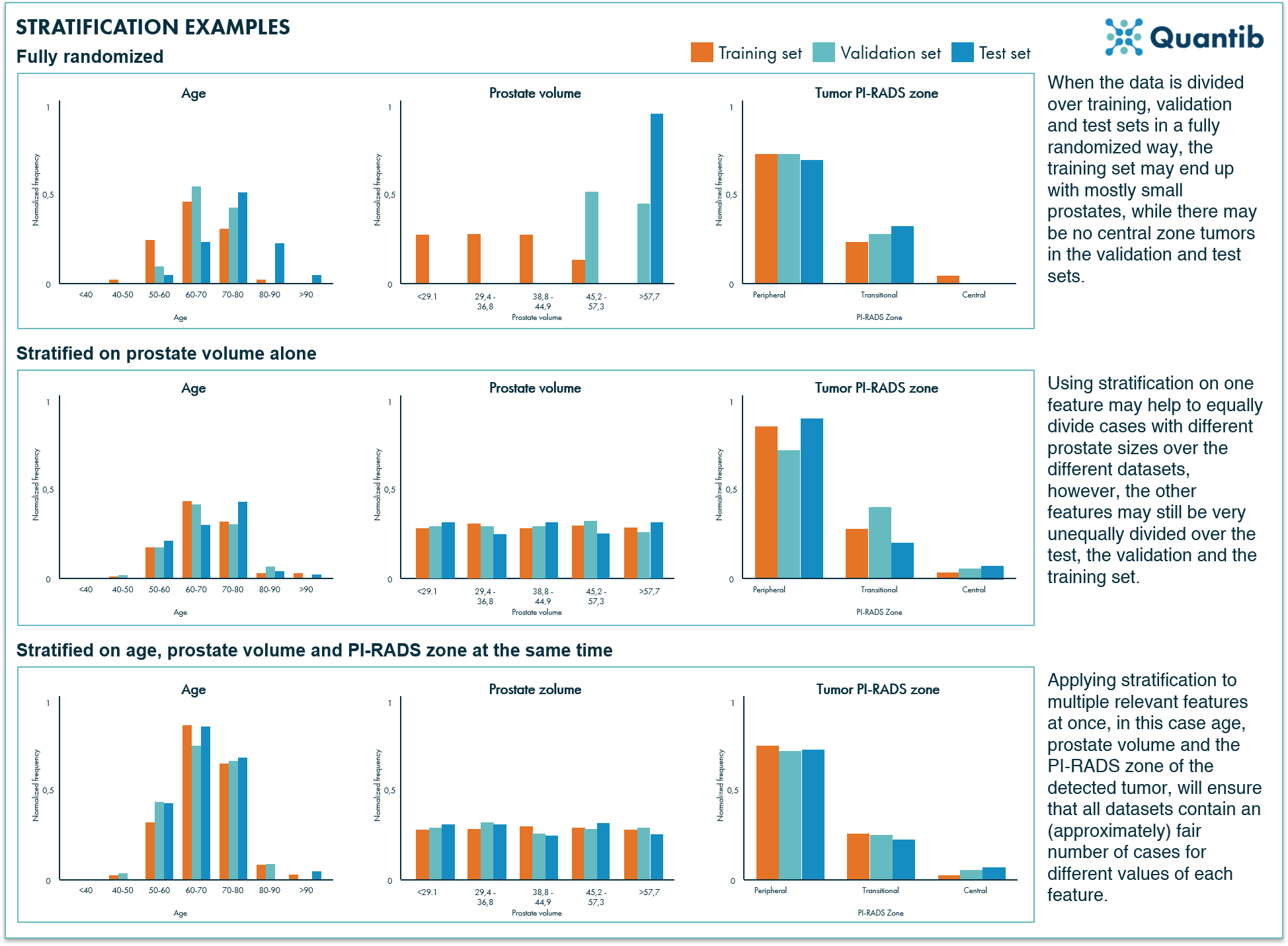
Figure 3: Examples of three different scenarios for data distribution over the different datasets. The fully randomized approach will most likely provide an unfair distribution of different values per feature, stratifying the data on one feature will improve the situation a little bit, however, the best way to go is applying stratification on all relevant features.
In conclusion
Medical datasets are usually small. Therefore, it is smart, if not crucial, to leverage the data that is available in an optimal way. One of the ways to increase the value of a medical dataset and get the most out of it, is by applying stratification when dividing the data into the three datasets needed for training, validation and testing. This technique can boost algorithm performance and has the potential to provide a better idea of how the algorithm will perform in the clinic. It goes without saying, however, that it is always better to have a larger dataset to start with.

