Organ volumes, tumor dimensions, size of lymph nodes - these are some examples of measures that provide valuable information for radiologists to use in diagnostic processes. However, obtaining those measurements manually is a tedious and time-consuming job.
Therefore, automatic medical image segmentation has been an important topic of research in the last decades. Artificial Intelligence (AI) algorithms are exquisitely suited to obtain measurements and segmentations from medical images, because they do not require hours of manual work. In addition, automatic analysis eliminates inter- and intra-observer differences, as the segmentation results will always be the same for the same input image. Deep learning-based methods have especially claimed some impressive results over the past few years. But how exactly do these algorithms work? And why are they so suitable for medical image segmentation?
In this article, we will explore what medical image segmentation is and the different segmentation algorithms currently available, starting from the classic approaches up to the latest trends in deep learning.
What is medical image segmentation?
Medical image segmentation consists of indicating the surface or volume of a specific anatomical structure in a medical image. Input images can range from X-rays and ultrasonography to CT and MRI scans. A segmentation algorithm will provide an output that indicates a region of interest as e.g. a contour or a label for every pixel or voxel in the image.
AI methods to segment medical images
Contour-based segmentation
A contour-based approach searches for the border between the structure to be segmented and its surroundings. This is quite similar to how a radiologist would manually segment an organ: by drawing a line between the organ and other body parts.
A classic contour-based segmentation method is the active contour model also known as snakes. This algorithm starts with an initial contour close to the object that you want to segment. Subsequently, it looks at voxels right next to those that are part of the contour to see whether these are brighter or darker than the contour voxels. Based on this information the contour shifts a little bit and checks again until it finds its optimal position. This optimal position depends on the provided rules like “move towards the brighter pixels”, or restrictions such as “avoid ending up with sharp corners”.
The difference with a voxel-based method is that it does not analyze every voxel in the image because the algorithm starts with a contour and adjusts the shape of this until it has found edges.
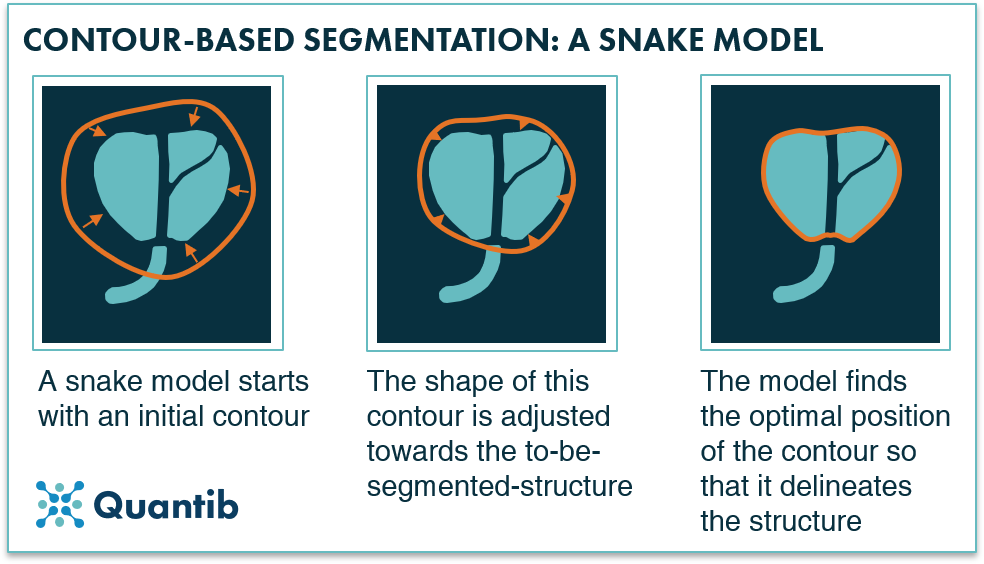
Figure 1: A snake model adjusts a contour in multiple steps to get to a delineation, a segmentation, of a structure.
Voxel-based segmentation
A voxel-based approach works in a similar way to classification methods because the algorithm looks at each voxel separately and determine whether it is part of the structure you want to segment. After addressing each voxel in the image, the algorithm will output a segmentation.
A very basic example of a voxel-based segmentation algorithm is simple thresholding. This method looks at the intensity of a voxel and compares it to a set threshold. If the voxel is above the value, the algorithm will mark it as part of the structure while if the voxel is below the value it will be marked as “surroundings” (or the other way around in case of hypo-intense structures). This method however only works well for structures that have a very different intensity from their surroundings and are not affected too much by noise.
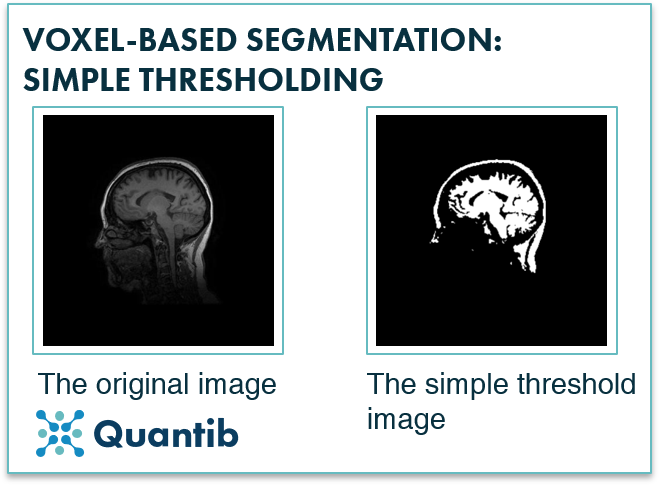

Figure 2: Simple thresholding is used to segment an image based on voxel intensity. As is already apparent from the image, it is tricky to segment only one structure with this approach as structures are rarely the only objects in an image that have an intensity above or below a certain value.
Registration-based methods
The general idea of a registration-based segmentation method is that you already have a segmented structure in one image (or a set of images), and you want to have this same structure segmented in a second image. You’ll need a transformation, meaning a very exact description of how you need to deform the first image, so that it becomes the second image. Applying this same transformation to the segmentation of the first image will provide you the segmentation for the second image.
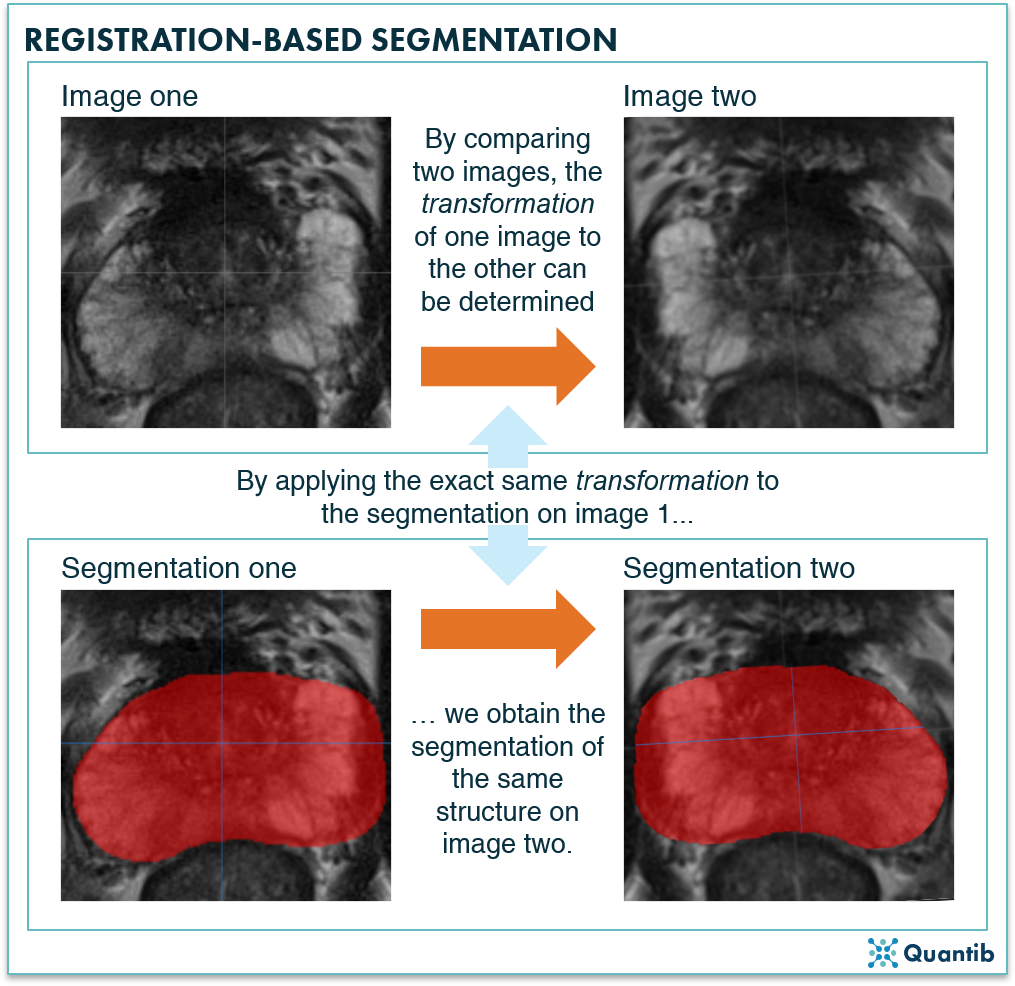
Figure 3: A registration-based segmentation algorithm first determines what adjustment, or transformation, is needed to change image one into image two. Secondly, applying this exact same transformation to an already available segmentation on image one, will lead to a segmentation of the same structure in image two.
Why use deep learning for image segmentation in radiology?
Over the past years deep learning has proven to be very suitable for the task, because of its great capabilities of adapting to a specific segmentation task by learning from example segmentations.
Which deep learning networks are the best for segmentation in radiology and how do they work?
Convolutional neural networks (CNNs)
A CNN is a specific form of a deep neural network. It uses a combination of convolution steps, meaning applying a certain filter to each pixel in the image, and pooling steps, meaning downsampling the image. These steps are performed in different layers of the neural network.
After applying both steps a couple of times, the algorithm has filtered out the most important information in the image and is able to determine what the image contains.
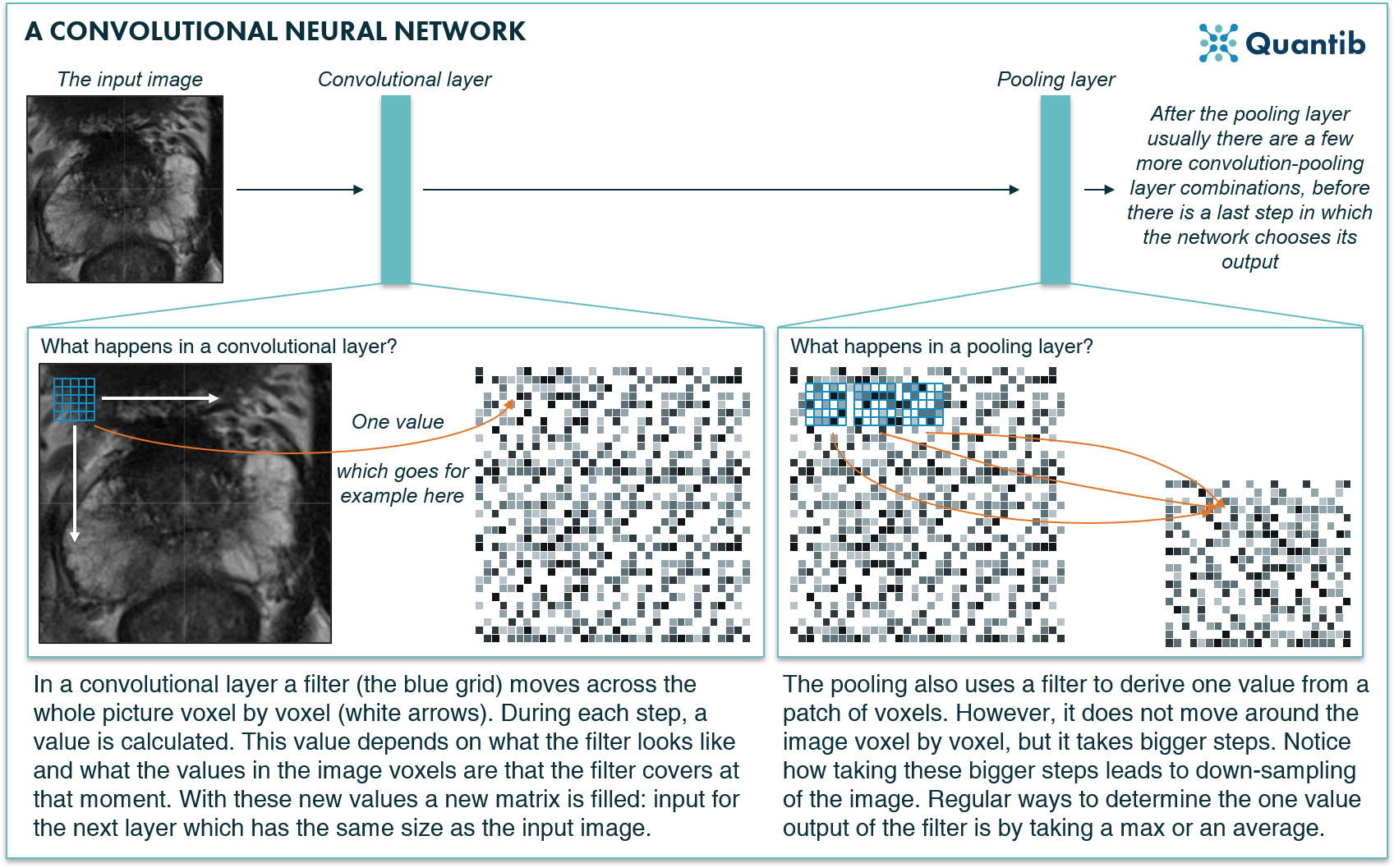
Figure 4: A convolutional neural network applies special convolution filters to a medical image to deduce the structures or organs present in the image.
Not every CNN is explicitly suited for segmenting objects. For (medical) image segmentation, a straightforward approach would be to apply a standard CNN to each voxel separately. For that, the CNN receives a selected patch around the voxel as input that will be classified. To segment the full image, this process will have to be repeated with all the voxels of the image.
Fully convolutional networks (FCNs)
An FCN can contain convolutional and pooling layers. However, it compensates for the downsampling in the pooling layers by adding upsampling layers that increase the dimensions of the image until it is at its original size, and can therefore be trained to produce a voxel-based segmentation as output1.
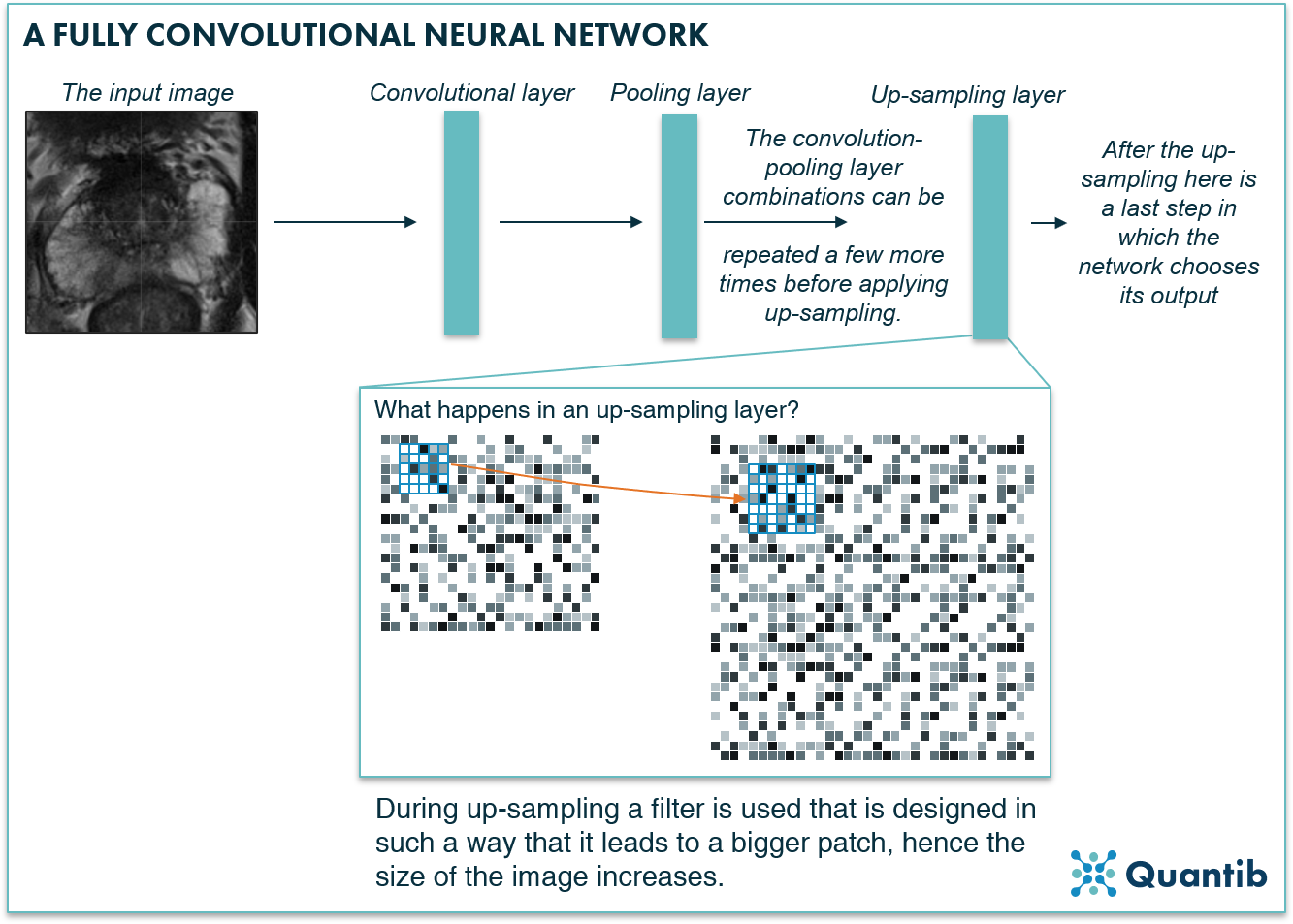
Figure 5: A fully convolutional neural network assures the segmentation output of the network has the same size as the input medical image by using upsampling layers.
U-net
Another example of a CNN that has the dedicated characteristics for image segmentation is a U-net. It contains both convolutional layers and pooling layers in the first part (the left side of the U in the image below), hence this is where the image gets downsampled. However, the network will upsample the image again by using upsampling layers until dimensions are the same as the input layer (the left side of the U in the image below). A U-net is very similar to an FCN, and you could even say it is a type of FCN. The characteristic factor is that a U-net has connections between the downward path and the upward path (grey arrows in the image below). Thus, it uses information in the upsampling process that was otherwise lost during downsampling2.
(3D) U-nets have been widely used for medical image segmentations in recent years. A very successful iteration is called nnU-net3 (no new net), which has shown great results in a large variety of segmentation tasks.
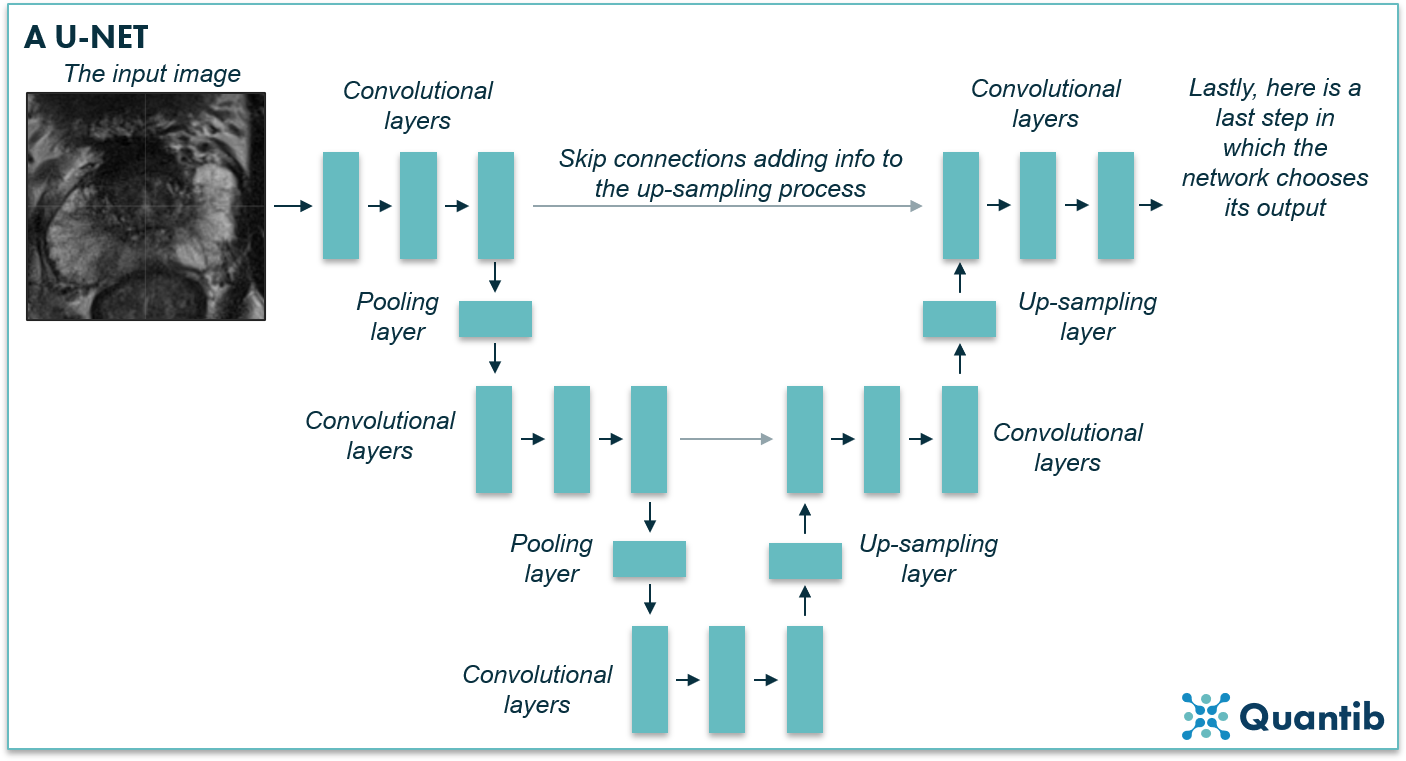
Figure 6: A U-net is a specific type of fully convolutional network, named after the U that is present in most schematic representations of such a network (just as in the image above).
Dilated convolutional network
Another type of network specifically developed for segmentation are dilated CNNs. Once you understand the basic idea of CNNs, the adjustment to get to dilated CNNs is a small step to grasp. Where regular CNNs use a filter of voxels that are all next to each other, dilated CCNs use a more “spread” filter, hence the filter uses information from voxels that are not direct neighbors, but that are spread over a wider area. What are the benefits of this method? Dilated CNNs use multiple layers of these special filters with different “dilation”, i.e. the distance between the voxels used in the filter is not always the same. This allows the network to include a lot of surrounding information to get to a decision for all voxels in the image without the use of pooling layers, hence the network does not downsample. So it also does not need to upsample to get to the right size for the output segmentation. Think of it as a similar approach to how a radiologist would look at a larger part of the image to recognize the anatomy4.
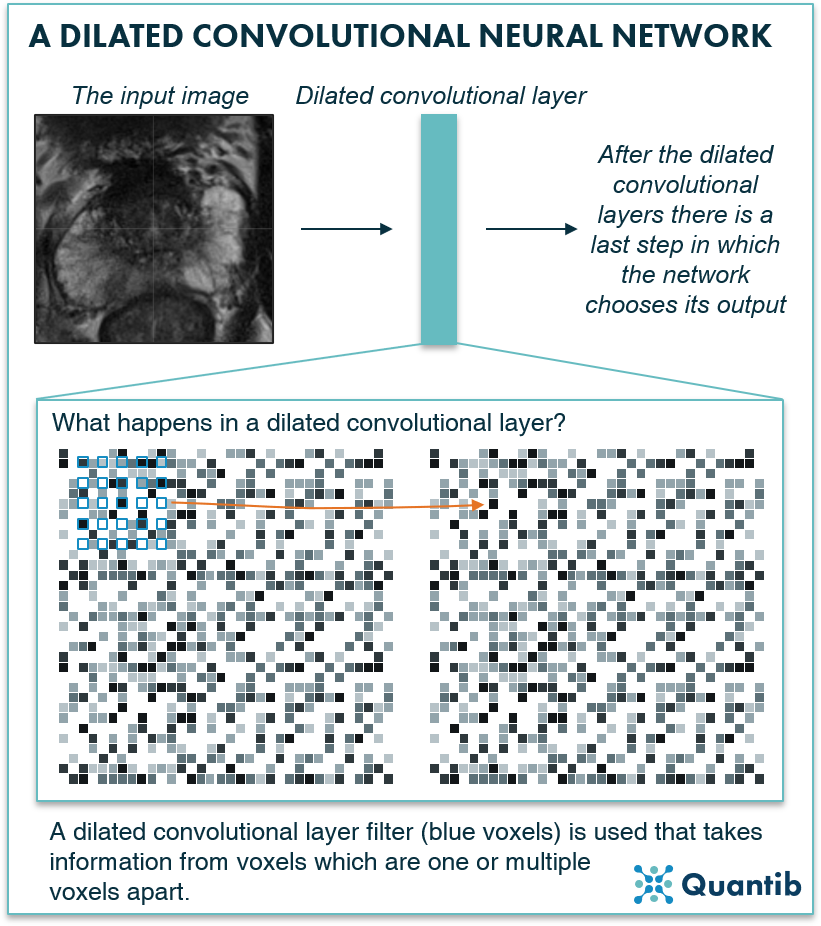
Figure 7: A dilated convolutional neural network uses dilation to collect information from a wider patch of pixels without using pooling layers, hence no upsampling is needed.
Transformers
In the recent years, transformers5 have been gaining popularity in various deep learning tasks, such as classification. Originally transformers are developed for natural language processing (NLP), where they take a sentence as input and translate such a sequence of words into another language. This approach can be used in computer vision tasks by splitting the image into a sequence of image patches6, corresponding to the sequence of words in NLP. In medical image segmentation this approach has been combined into U-net like networks such as TransUNet7 and UNETR8, showing promising results.
Conclusion
As you can see, there are many different possibilities to approach medical image segmentation. None are necessarily “better” than the others, because this very much depends on your training data, input scans, and what you are trying to segment (to name a few factors). Deep learning, and especially FCNs, are exquisitely suited for image segmentation. Many research groups have used such networks, or variations on those, for medical segmentation purposes. Results have been impressive, but this does not mean that they can directly be applied in a clinical setting.
Discover Quantib's FDA cleared AI-driven software for prostate MRI support and brain atrophy quantification.
Bibliography
-
Long, J., Shelhamer, E. & Darrell, T. Fully convolutional networks for semantic segmentation. In 2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 3431–3440 (IEEE, 2015). doi:10.1109/CVPR.2015.7298965.
-
Ronneberger, O., Fischer, P. & Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. (MICCAI, 2015).
-
Isensee, F., Jaeger, P. F., Kohl, S. A. A., Petersen, J. & Maier-Hein, K. H. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat Methods 18, 203–211 (2021).
-
Yu, F. & Koltun, V. Multi-Scale Context Aggregation by Dilated Convolutions. (ICLR, 2016).
-
Vaswani, A. et al. Attention is All you Need. in Advances in Neural Information Processing Systems (eds. Guyon, I. et al.) vol. 30 (Curran Associates, Inc., 2017).
-
Dosovitskiy, A. et al. An Image is Worth 16x16 Words: Transformers for Image Recognition at Scale. (ICLR, 2021).
-
Chen, J. et al. TransUNet: Transformers Make Strong Encoders for Medical Image Segmentation. (2021).
-
Hatamizadeh, A. et al. UNETR: Transformers for 3D Medical Image Segmentation. (WACV, 2022).